- Share
- Share on Facebook
- Share on LinkedIn
AMP-activated protein kinase (AMPK)
AMP-activated protein kinase (AMPK) is an evolutionary conserved heterotrimeric complex that emerged as the central hub for cellular energy signaling. AMPK is capable of sensing and responding to changes in cellular energy state. Its complex activation mechanism comprises allosteric activation by a failing energy state (low ATP/ADP and ATP/AMP ratios) and covalent activation controlled by calcium and extracellular signals. However, many underlying molecular mechanisms remain elusive.
We first described that binding of AMP (and ADP) induces a conformational switch within AMPK as a central part of the activation mechanism (1). This was later confirmed by X-ray crystallography and other studies (2). Based on this knowledge, we engineered a first genetically encoded, synthetic fluorescent nano-sensor for the analysis of allosteric AMPK activation and the real-time tracking of energy state in single cells (3). This technology holds promise for drug screening and as a readout for stress and cellular health.
Activation of AMPK also differs in different tissues and organs. The heart particularly relies on constant energy supply, but the role of cardiac AMPK is incompletely understood. We are studying the first cardiac mouse model where both catalytic AMPK subunits can be invalided conditionally at adult age.
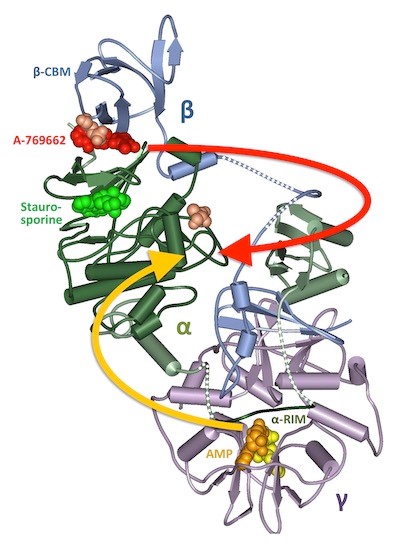
Viollet et al. (2014) Chemistry & Biology 21, 567-9.
NME4 protein
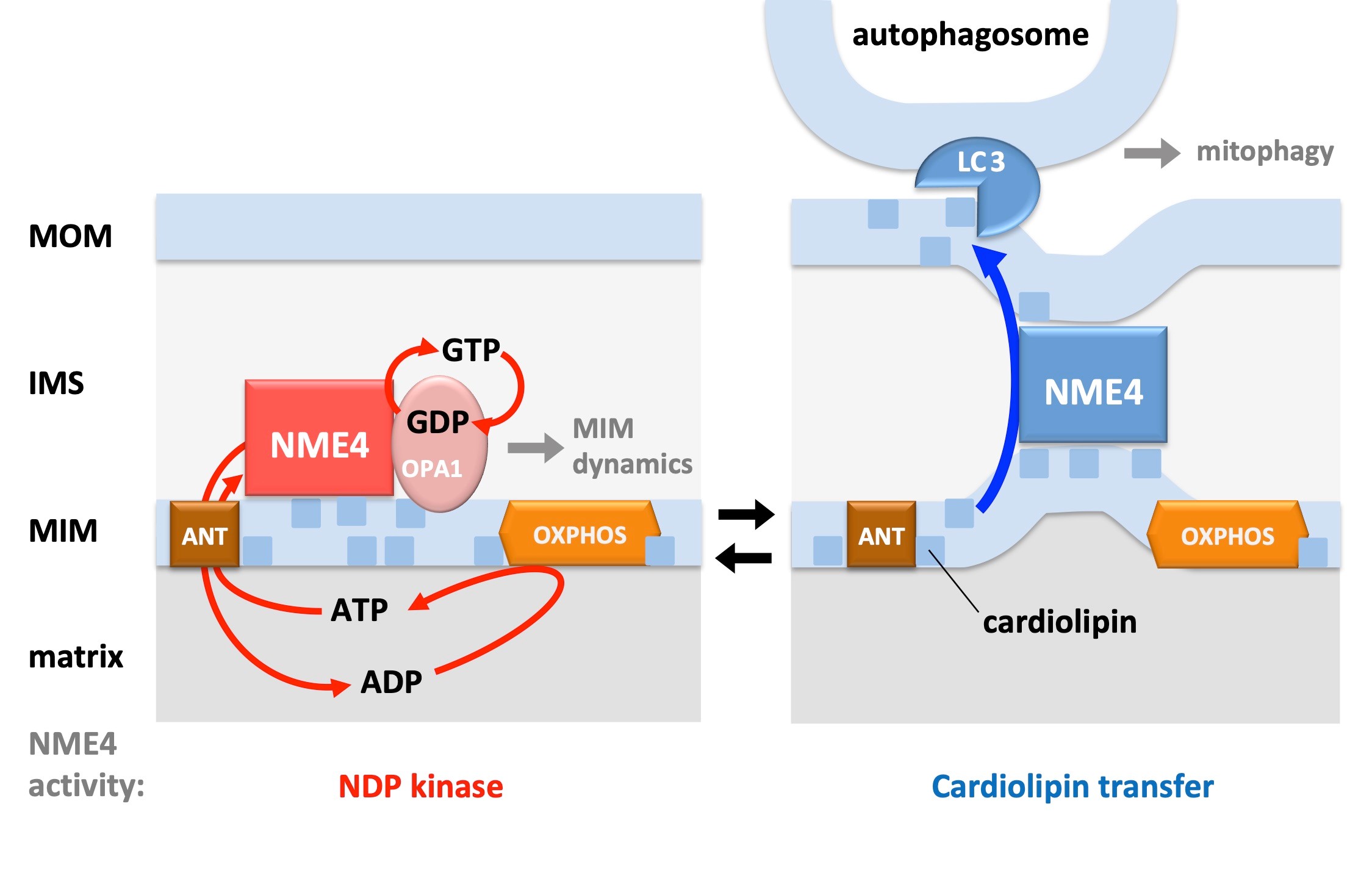
NME4 belongs to a mammalian family of multi-functional proteins. A basic property of many members is NDP kinase activity, i.e. the use of ATP to generate other NTPs. We have shown that NME4 is specifically localized in the mitochondrial intermembrane space (4), where it interacts with the dynamin-related GTPase OPA1 (5) to fuel it locally with the necessary GTP (6). More recently, we evidenced an original role of NME4 in the transfer of the mitochondrial phospholipid cardiolipin from the inner membrane to the mitochondrial surface (5,7). There, cardiolipin can acts as a signal triggering mitophagy or apoptosis. We are now studying the consequences of NME4 loss-of-function and knock-out at the cellular and organism levels, respectively.
Recently, we have organized the 11th international conference on NME proteins.
Creatine kinase
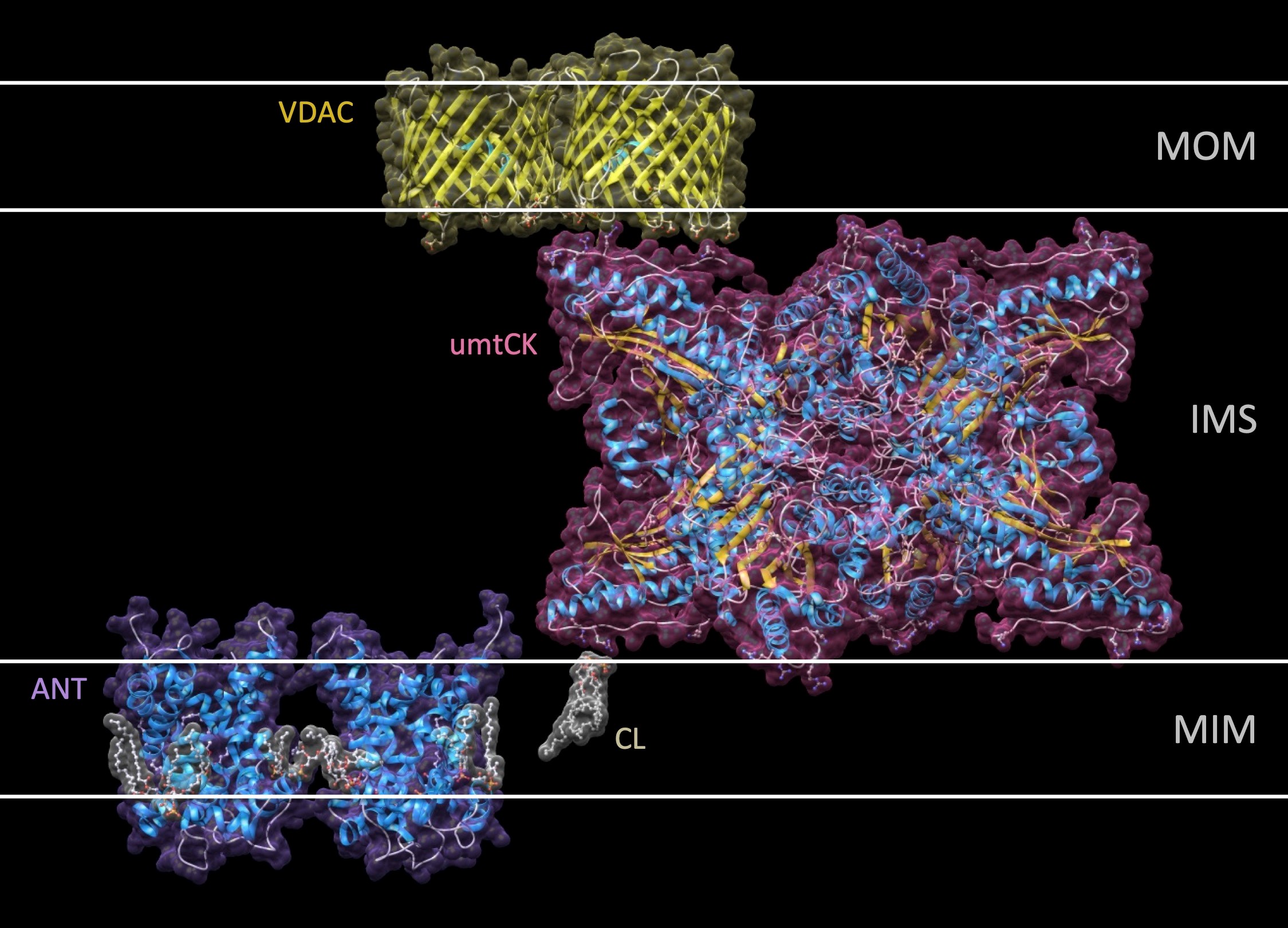
Isoforms of creatine kinase (CK) are central to an intricate cellular energy buffering and transfer system that maintains energy homeostasis in cells with high or fluctuating energy requirements. After having solved the molecular structure of different CKs, we have been interested in characterizing the molecular physiology, in particular of mitochondrial CK isoforms (reviewed in (8,9)).
References
(1) Riek, U., Scholz, R., Konarev, P. Rufer, A., Suter, M., Nazabal, A., Ringler, P., Chami, M., Müller, S.A., Neumann, D., Forstner, M. Hennig, M., Zenobi, R., Engel, A. Svergun, D., Schlattner, U.*, and Wallimann, T.* (2008) Structural properties of AMP-activated protein kinase (AMPK) : dimerization, molecular shape, and changes upon ligand binding. J. Biol. Chem. 283, 18331-18343.
(2) Chen, L., Wang, J., Zhang, Y.Y., Yan, S.F., Neumann, D., Schlattner, U., Wang, Z.X., and Wu, J.W. (2012) AMP-activated protein kinase undergoes nucleotide-dependent conformational changes. Nature Struct. Mol. Biol. 19, 716-8.
(3) Pelosse, M., Cottet-Rousselle, C., Bidan, C., Dupont, A., Gupta, K., Berger, I., and Schlattner, U. (2019) Synthetic energy sensor AMPfret deciphers adenylate-dependent AMPK activation mechanism. Nat. Commun. 10, 1-13.
(4) Tokarska-Schlattner, M., Boissan, M., Munier, A., Borot, C., Mailleau, C., Speer, O., Schlattner, U.*, and Lacombe, M.-L.* (2008) The nucleoside diphosphate kinase D (NM23-H4) binds the mitochondrial inner membrane with high affinity to cardiolipin and couples nucleotide transfer with respiration. J. Biol. Chem. 283, 26198-207.
(5) Schlattner, U., Tokarska-Schlattner, M., Ramirez Rios, S., Seffouh, A., Tyuina, Y.Y., Amoscato, A.A., Huang, Z., Jiang, J., Boissan, M., Epand, R.F., Mohammadsanyi, D., Klein-Seetharaman, J., Epand, R.M., Lacombe, M.L., and Kagan, V.E. (2013) Dual function of mitochondrial Nm23-H4 protein in phosphotransfer and intermembrane lipid transfer: a cardiolipin switch. J. Biol. Chem. 288, 111-121.
(6) Boissan, M., Montagnac, G., Shen, Q., Griparic, L., Guitton, J., Romao, M., Sauvonnet, N., Lagache, T., Lascu, I., Raposo, G.,Desbourdes, C., Schlattner, U., Lacombe, M.L., Polo, S., van der Bliek, A.M., Roux, A., and Chavrier, P. (2014) Nucleoside diphosphate kinases fuel dynamin superfamily proteins with GTP for membrane remodeling. Science 344, 1510-5.
(7) Kagan, E., Jiang, J., Huang, Z., Tyurina, Y.Y., Desbourdes, C., Cottet-Rousselle, C., Dar, H.H., Verma, M., Tyurin, V.A., Kapralov, A.A., Cheikhi, A., Mao, G., Stolz, D., St. Croix, C.M., Watkins, S., Shen, Z., Li, Y., Greenberg, M.L., Tokarska-Schlattner, M., Boissan, M., Lacombe, M.L., Epand, R.M., Chu, C.T., Mallampalli, R., Bayir, H., and Schlattner, U. (2016) NDPK-D (NM23-H4)-mediated externalization of cardiolipin enables elimination of depolarized mitochondria by mitophagy. Cell Death Diff. 23, 1140-51
(8) Schlattner, U., Tokarska-Schlattner, M., and Wallimann, T. (2006) Mitochondrial creatine kinase in human health and disease. Biochem. Biophys. Acta 1762, 164-180.
(9) Schlattner, U., Kay, L., and Tokarska-Schlattner, M. (2018) Mitochondrial proteolipid complexes of creatine kinase. Subcell. Biochem. 87, 365-408.
Biophysical Society – Bioenergetics, Mitochondria and Metabolism Subgroup
Uwe Schlattner, co-chair of the subgroup, provides here the presentation of the 2022 business meeting in San Francisco.
Read the presentation
Coordinator

Uwe Schlattner, PU
uwe.schlattner univ-grenoble-alpes.fr (uwe[dot]schlattner[at]univ-grenoble-alpes[dot]fr)
univ-grenoble-alpes.fr (uwe[dot]schlattner[at]univ-grenoble-alpes[dot]fr)
Phone number: +33 (0)4 76 51 46 71
- Share
- Share on Facebook
- Share on LinkedIn